How to use Octave to Fit an Arbitrary Function with fmins: Difference between revisions
No edit summary |
No edit summary |
||
| (5 intermediate revisions by the same user not shown) | |||
| Line 1: | Line 1: | ||
Suppose we want to fit experimental data to an arbitrary function. For our purposes let that function be <math>y = A cos(t + \theta) </math>. We can use the fmins function from the octave package [http://octave.sourceforge.net/optim/index.html optim]. We are looking for a best fit <math>\theta</math> and <math>A</math>. The fmins function will try a whole lot of different values for these parameters until it decides to give up of it has found a local minimum. |
Suppose we want to fit experimental data to an arbitrary function. For our purposes let that function be <math>y = A cos(t + \theta) </math>. We can use the fmins function from the octave package [http://octave.sourceforge.net/optim/index.html optim]. We are looking for a best fit <math>\theta</math> and <math>A</math>. The fmins function will try a whole lot of different values for these parameters until it decides to give up of it has found a local minimum. |
||
The first step is to construct a function that computes the sum of the differences between the guess for the best fit function and the experimental data. This is the sum of the squared errors at each data point. The idea is that octave will use the fmins function to find the parameters that minimize this sum of squared errors. This |
The first step is to construct a function that computes the sum of the differences between the guess for the best fit function and the experimental data. This is the sum of the squared errors at each data point. The idea is that octave will use the fmins function to find the parameters that minimize this sum of squared errors. This needs to be put in a separate dot m file called model.m (the same as the function). |
||
function sum_square_errors = model(p,x,y) |
function sum_square_errors = model(p,x,y) |
||
| Line 11: | Line 11: | ||
difference=y-y_trial; % The difference between fit and experimental data |
difference=y-y_trial; % The difference between fit and experimental data |
||
sum_square_errors = sum(difference.^2); % The sum of the squared errors |
sum_square_errors = sum(difference.^2); % The sum of the squared errors |
||
You can put the above in a separate file called "sum_square_errors.m", or put it in your main file. |
|||
The second step is to call fmins with the appropriate arguments to minimize the sum of the squared errors. For this example I will make up some data, add noise to it and call it y. In real life, you will probably type your vectors of x and y in by hand. |
The second step is to make a script to call fmins with the appropriate arguments to minimize the sum of the squared errors. For this example I will make up some data, add noise to it and call it y. In real life, you will probably type your vectors of x and y in by hand. |
||
x=0:.1:2*pi; |
x=0:.1:2*pi; |
||
| Line 20: | Line 18: | ||
y_data=y_true+randn(size(x)); % Add noise to our true values |
y_data=y_true+randn(size(x)); % Add noise to our true values |
||
p0=[1.5 1] % Guessed parameters. |
p0=[1.5 1] % Guessed parameters. |
||
p=fmins( |
p=fmins('model', p0, [],[], x, y_data); % Use fmins to find the best fit p |
||
plot(x,y_true,x,y_data,'r*',x,p(1)*cos(x+p(2)),'g-') |
plot(x,y_true,x,y_data,'r*',x,p(1)*cos(x+p(2)),'g-') |
||
legend('True', 'Noisy', 'Best Fit') |
legend('True', 'Noisy', 'Best Fit') |
||
title('Fitting Example with fmins') |
title('Fitting Example with fmins') |
||
xlabel('x') |
xlabel('x') |
||
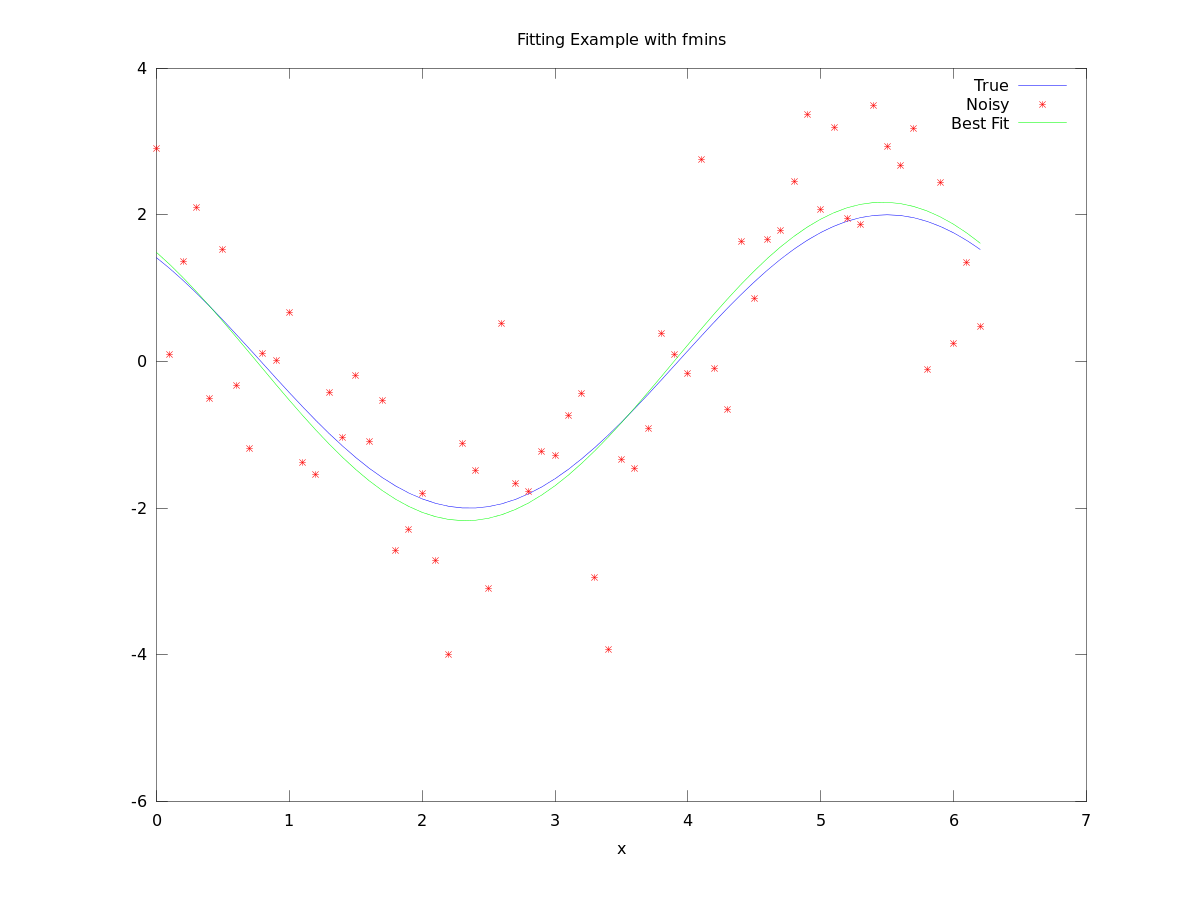
[[File:Fit_example.png]] |
|||
function sse = expmodel(p,x,y) |
|||
%EXPMODEL Fit of a 2-term exponential model to a set of data |
|||
% The parameters are in p (4-by-1 vector) and assumed to |
|||
% be in the following order: |
|||
a1 = p(1); |
|||
b1 = p(2); |
|||
a2 = p(3); |
|||
b2 = p(4); |
|||
ypred = a1*exp(b1*x) - a2*exp(b2*x); % predictions |
|||
err = y - ypred; % mismatches |
|||
sse = sum(err.^2); % residual measure |
|||
Latest revision as of 18:11, 6 January 2011
Suppose we want to fit experimental data to an arbitrary function. For our purposes let that function be . We can use the fmins function from the octave package optim. We are looking for a best fit and . The fmins function will try a whole lot of different values for these parameters until it decides to give up of it has found a local minimum.
The first step is to construct a function that computes the sum of the differences between the guess for the best fit function and the experimental data. This is the sum of the squared errors at each data point. The idea is that octave will use the fmins function to find the parameters that minimize this sum of squared errors. This needs to be put in a separate dot m file called model.m (the same as the function).
function sum_square_errors = model(p,x,y) % This is where we computer the sum of the square of the errors. % The parameters are in the vector p, which for us is a two by one. A=p(1); theta=p(2); y_trial=A*cos(x+theta); % y with the trial parameters. difference=y-y_trial; % The difference between fit and experimental data sum_square_errors = sum(difference.^2); % The sum of the squared errors
The second step is to make a script to call fmins with the appropriate arguments to minimize the sum of the squared errors. For this example I will make up some data, add noise to it and call it y. In real life, you will probably type your vectors of x and y in by hand.
x=0:.1:2*pi;
y_true = 2*cos(x+pi/4);
y_data=y_true+randn(size(x)); % Add noise to our true values
p0=[1.5 1] % Guessed parameters.
p=fmins('model', p0, [],[], x, y_data); % Use fmins to find the best fit p
plot(x,y_true,x,y_data,'r*',x,p(1)*cos(x+p(2)),'g-')
legend('True', 'Noisy', 'Best Fit')
title('Fitting Example with fmins')
xlabel('x')